SUMO Antibody
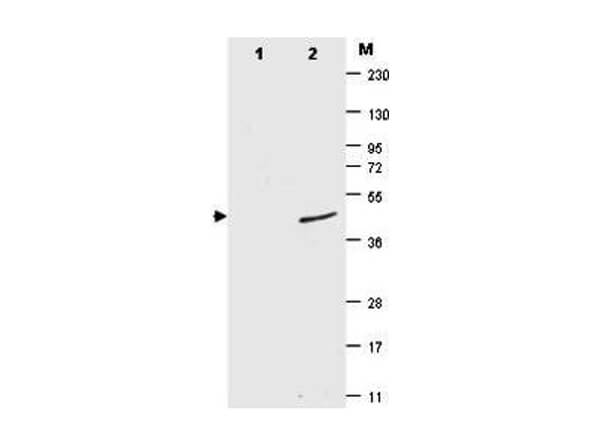
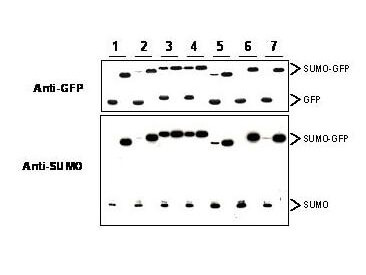
Mouse Monoclonal 4F2.F5.G2 IgG1 kappa
$50.00 to US & $70.00 to Canada for most products. Final costs are calculated at checkout.
Background
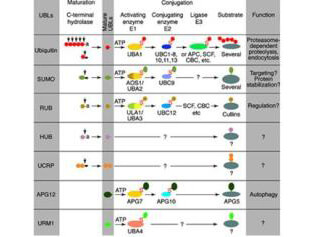
Covalent modification of cellular proteins by the ubiquitin-like modifier SUMO (small ubiquitin-like modifier) regulates various cellular processes, such as nuclear transport, signal transduction, stress responses and cell cycle progression. But, in contrast to ubiquination, sumoylation does not tag proteins for degradation by the 26S proteasome, but rather seems to enhance stability or modulate their subcellular compartmentalization. Ubiquitin-like proteins fall into two classes: the first class, ubiquitin-like modifiers (UBLs) function as modifiers in a manner analogous to that of ubiquitin. Examples of UBLs are SUMO, Rub1 (also called Nedd8), Apg8 and Apg12. Proteins of the second class include parkin, RAD23 and DSK2, are designated ubiquitin-domain proteins (UDPs). These proteins contain domains that are related to ubiquitin but are otherwise unrelated to each other. In contrast to UBLs, UDPs are not conjugated to other proteins. Once covalently attached to cellular targets, SUMO regulates protein:protein and protein:DNA interactions, as well as localization and stability of the target protein. Sumoylation occurs in most eukaryotic systems, and SUMO is highly conserved from yeast to humans. Where invertebrates have only a single SUMO gene termed SMT3, three members of the SUMO family have been identified in vertebrates: SUMO-1 and the close homologues SUMO-2 and SUMO-3. SUMO has been called SMT3 (yeast), sentrin, PIC1, GMP1 and UBL1. SUMO has been shown to bind and regulate mammalian SP-RINGs (such as Mdm2, PIAS and PML), RanGAP1, RanBP2, p53, p73, HIPK2, TEL, c-Jun, Fas, Daxx, TNFRI, Topo-I, Topo-II, WRN, Sp100, IkB-a, Androgen receptor (AR), GLUT1/4, Drosophila Ttk69, Dorsal, CaMK, yeast Septins, and viral CMV-IE1/2, EBV-BZLF1, HPV/BPV-E1. These bindings implicate SUMO in the stabilization of the target proteins and/or their localization to subcellular complexes. SUMO has an apparent molecular weight of ~12kDa and human SUMO-1 (a 101 amino acid polypeptide) shares 50% sequence identity with SUMO-2 and SUMO-3 and with yeast SMT3. SUMO and ubiquitin only show about 18% homology, but both possess a common three-dimensional structure characterized by a tightly packed globular fold with b-sheets wrapped around an a-helix.
Product Details
Target Details
Application Details
Formulation
Shipping & Handling
This product is for research use only and is not intended for therapeutic or diagnostic applications. Please contact a technical service representative for more information. All products of animal origin manufactured by Rockland Immunochemicals are derived from starting materials of North American origin. Collection was performed in United States Department of Agriculture (USDA) inspected facilities and all materials have been inspected and certified to be free of disease and suitable for exportation. All properties listed are typical characteristics and are not specifications. All suggestions and data are offered in good faith but without guarantee as conditions and methods of use of our products are beyond our control. All claims must be made within 30 days following the date of delivery. The prospective user must determine the suitability of our materials before adopting them on a commercial scale. Suggested uses of our products are not recommendations to use our products in violation of any patent or as a license under any patent of Rockland Immunochemicals, Inc. If you require a commercial license to use this material and do not have one, then return this material, unopened to: Rockland Inc., P.O. BOX 5199, Limerick, Pennsylvania, USA.